In this tutorial we will reconstruct a 2D MR image from multicoil Cartesian under-sampled kspace measurements.
We use the toy datasets available in pysap, more specifically a 2D brain slice and under-sampled Cartesian acquisition over 32 channels. We compare zero-order image reconstruction with calibrationless multi-coil Compressed sensing reconstructions (analysis vs synthesis formulation) using the FISTA algorithm for the synthesis formulation and the Condat-Vu algorithm for the analysis formulation. Structured sparsity will be promoted in the wavelet domain, using either Symmlet-8 (analysis and synthesis) or undecimated bi-orthogonal wavelets (analysis only) considering group-LASSO or OSCAR-based regularization. The multicoil data is collected across multiple, say , channels.
We remind that the synthesis formulation of the Calibrationless CS-PMRI problem reads (minimization in the sparsifying domain):
where and such that . The image solution is given by . For an orthonormal wavelet transform, we have while for a frame we may have . The regularization term promotes structured sparsity. For instance when one chooses group-LASSO regularization , where the L2 norm involves the channels per wavelet coefficient .
The analysis formulation consists in minimizing the following cost function (min. in the image domain):
Author: Chaithya G R & Philippe Ciuciu
Date: 01/07/2021
Target: ATSI MSc students, Paris-Saclay University
# Package import
from mri.operators import FFT, WaveletN, OWL
from mri.reconstructors import CalibrationlessReconstructor
from pysap.data import get_sample_data
# Third party import
from modopt.opt.proximity import GroupLASSO
from modopt.math.metrics import ssim
import numpy as np
import matplotlib.pyplot as pltWARNING: Using pyFFTW "monkey patch" for scipy.fftpack
# Loading input data
cartesian_ref_image = get_sample_data('2d-pmri').data
image = np.linalg.norm(cartesian_ref_image, axis=0)
# Obtain MRI cartesian mask
mask = get_sample_data("cartesian-mri-mask").data# View Input
plt.subplot(1, 2, 1)
plt.imshow(np.abs(image), cmap='gray')
plt.title("MRI Data")
plt.subplot(1, 2, 2)
plt.imshow(mask, cmap='gray')
plt.title("K-space Sampling Mask")
plt.show()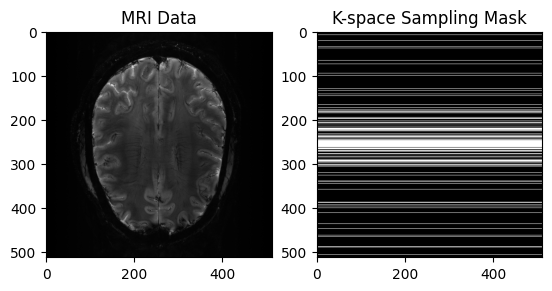
Generate the kspace¶
From the 2D brain slice and the acquisition mask, we retrospectively undersample the k-space using a cartesian acquisition mask We then reconstruct the zero order solution as a baseline
# Get the locations of the kspace samples and the associated observations
fourier_op = FFT(mask=mask, shape=image.shape,
n_coils=cartesian_ref_image.shape[0])
kspace_obs = fourier_op.op(cartesian_ref_image)# Zero order solution
zero_soln = np.linalg.norm(fourier_op.adj_op(kspace_obs), axis=0)
base_ssim = ssim(zero_soln, image)
plt.imshow(np.abs(zero_soln), cmap='gray')
plt.title('Zero Order Solution : SSIM = ' + str(np.around(base_ssim, 3)))
plt.show()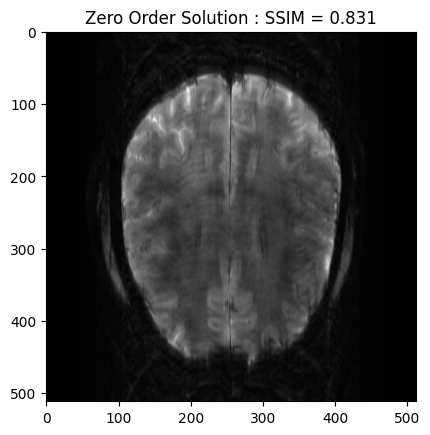
Synthesis formulation: FISTA vs POGM optimization¶
We now want to refine the zero order solution using a FISTA optimization. The cost function is set to Proximity Cost + Gradient Cost
# Setup the operators
linear_op = WaveletN(
wavelet_name='sym8',
nb_scale=4,
n_coils=cartesian_ref_image.shape[0],
)
coeffs = linear_op.op(cartesian_ref_image)
regularizer_op = GroupLASSO(weights=6e-8)Setup reconstructor:¶
# Setup Reconstructor
reconstructor = CalibrationlessReconstructor(
fourier_op=fourier_op,
linear_op=linear_op,
regularizer_op=regularizer_op,
gradient_formulation='synthesis',
verbose=1,
)WARNING: Making input data immutable.
Lipschitz constant is 1.1
# Run the FISTA reconstruction and view results
image_rec, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='fista',
num_iterations=100,
)
image_rec = np.linalg.norm(image_rec, axis=0)
recon_ssim = ssim(image_rec, image)
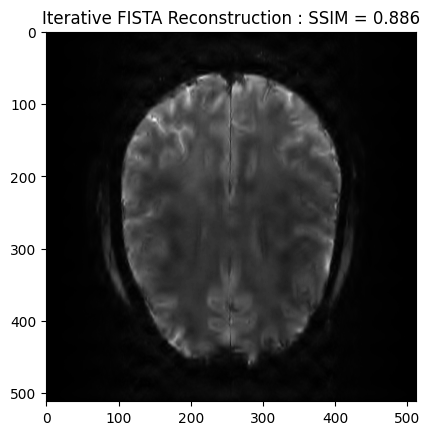
plt.imshow(np.abs(image_rec), cmap='gray')
plt.title('Iterative FISTA Reconstruction : SSIM = ' + str(np.around(recon_ssim, 3)))
plt.show()WARNING: Making input data immutable.
- mu: 6e-08
- lipschitz constant: 1.1
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x795b2c947df0> - 4
- max iterations: 100
- image variable shape: (512, 512)
- alpha variable shape: (32, 291721)
----------------------------------------
Starting optimization...
- final iteration number: 100
- final log10 cost value: 6.0
- converged: False
Done.
Execution time: 97.17409577900253 seconds
----------------------------------------
POGM optimization¶
# Run the POGM reconstruction and view results
image_rec2, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='pogm',
num_iterations=100,
)
image_rec2 = np.linalg.norm(image_rec2, axis=0)
recon2_ssim = ssim(image_rec2, image)
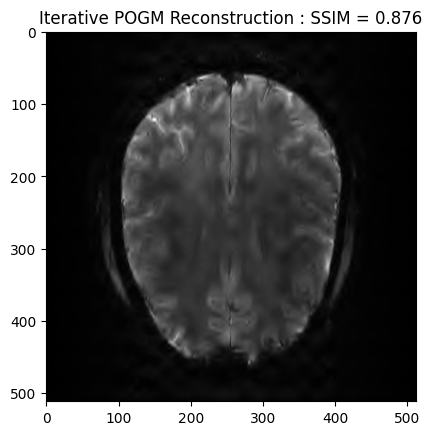
plt.imshow(np.abs(image_rec2), cmap='gray')
plt.title('Iterative POGM Reconstruction : SSIM = ' + str(np.around(recon2_ssim, 3)))
plt.show() - mu: 6e-08
- lipschitz constant: 1.1
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x795b2c947df0> - 4
- max iterations: 100
- image variable shape: (32, 512, 512)
----------------------------------------
Starting optimization...
- final iteration number: 100
- final log10 cost value: 6.0
- converged: False
Done.
Execution time: 105.65811157200005 seconds
----------------------------------------
# Setup the operators
linear_op = WaveletN(
wavelet_name='sym8',
nb_scale=4,
n_coils=cartesian_ref_image.shape[0],
)
coeffs = linear_op.op(cartesian_ref_image)
regularizer_op = OWL(
alpha=1.05e-8,
beta=0,
mode='band_based',
n_coils=cartesian_ref_image.shape[0],
bands_shape=linear_op.coeffs_shape,
)
# Setup Reconstructor
reconstructor = CalibrationlessReconstructor(
fourier_op=fourier_op,
linear_op=linear_op,
regularizer_op=regularizer_op,
gradient_formulation='synthesis',
verbose=1,
)WARNING: Making input data immutable.
Lipschitz constant is 1.0999998033046723
# Run the FISTA reconstruction and view results
image_rec, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='fista',
num_iterations=100,
)
image_rec = np.linalg.norm(image_rec, axis=0)
recon_ssim = ssim(image_rec, image)
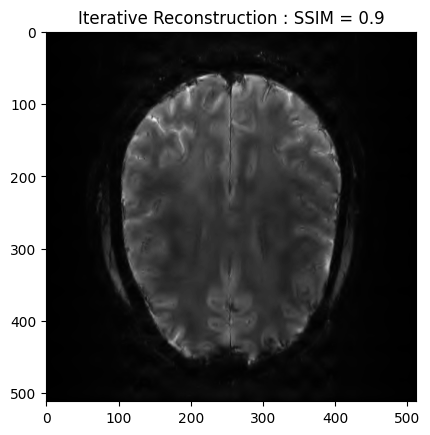
plt.imshow(np.abs(image_rec), cmap='gray')
plt.title('Iterative Reconstruction : SSIM = ' + str(np.around(recon_ssim, 2)))
plt.show() - mu: [<modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c87b6d0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c87b670>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c87bca0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b5d0be9e0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a140>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a350>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a0b0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a0e0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a440>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a410>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a020>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a260>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c83a680>]
- lipschitz constant: 1.0999998033046723
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x795b2c9479a0> - 4
- max iterations: 100
- image variable shape: (512, 512)
- alpha variable shape: (32, 291721)
----------------------------------------
Starting optimization...
- final iteration number: 100
- final log10 cost value: 6.0
- converged: False
Done.
Execution time: 195.9484668019977 seconds
----------------------------------------
linear_op = WaveletN(
wavelet_name='sym8',
nb_scale=4,
n_coils=cartesian_ref_image.shape[0],
)
regularizer_op = GroupLASSO(6e-8)reconstructor = CalibrationlessReconstructor(
fourier_op=fourier_op,
linear_op=linear_op,
regularizer_op=regularizer_op,
gradient_formulation='analysis',
verbose=1,
)WARNING: Making input data immutable.
Lipschitz constant is 1.0999999344348907
x_final, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='condatvu',
num_iterations=100,
)
image_rec = np.linalg.norm(x_final, axis=0)
recon_ssim = ssim(image_rec, image)
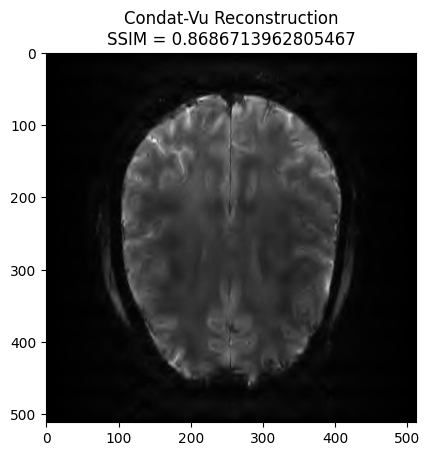
plt.imshow(np.abs(image_rec), cmap='gray')
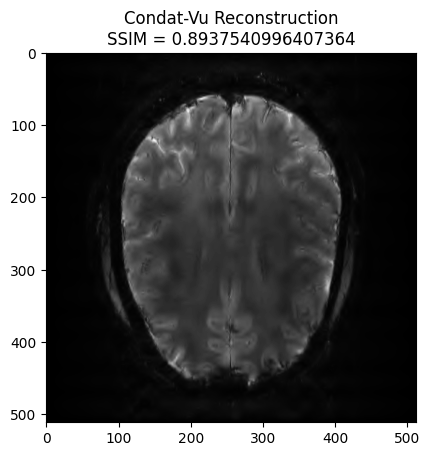
plt.title('Condat-Vu Reconstruction\nSSIM = ' + str(recon_ssim))
plt.show() - mu: 6e-08
- lipschitz constant: 1.0999999344348907
- tau: 0.9523809730454954
- sigma: 0.5
- rho: 1.0
- std: None
- 1/tau - sigma||L||^2 >= beta/2: True
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x795b2cd0d690> - 4
- max iterations: 100
- number of reweights: 0
- primal variable shape: (32, 512, 512)
- dual variable shape: (32, 291721)
----------------------------------------
Starting optimization...
WARNING: <class 'mri.operators.linear.wavelet.WaveletN'> does not inherit an operator parent.
- final iteration number: 100
- final cost value: 1000000.0
- converged: False
Done.
Execution time: 98.07776652800021 seconds
----------------------------------------
coeffs = linear_op.op(cartesian_ref_image)
regularizer_op = OWL(
alpha=1.05e-8,
beta=0,
mode='band_based',
n_coils=cartesian_ref_image.shape[0],
bands_shape=linear_op.coeffs_shape,
)reconstructor = CalibrationlessReconstructor(
fourier_op=fourier_op,
linear_op=linear_op,
regularizer_op=regularizer_op,
gradient_formulation='analysis',
verbose=1,
)WARNING: Making input data immutable.
Lipschitz constant is 1.100000262260437
x_final, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='condatvu',
num_iterations=100,
)
image_rec = np.linalg.norm(x_final, axis=0)
recon_ssim = ssim(image_rec, image)
plt.imshow(np.abs(image_rec), cmap='gray')
plt.title('Condat-Vu Reconstruction\nSSIM = ' + str(recon_ssim))
plt.show() - mu: [<modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c772920>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c751660>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c7519f0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c99e020>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c99e5f0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2cd46c80>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c9e9930>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c9e96c0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c9eabc0>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c8eb460>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c8eb430>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c8eb220>, <modopt.opt.proximity.OrderedWeightedL1Norm object at 0x795b2c8ebf70>]
- lipschitz constant: 1.100000262260437
- tau: 0.952380824371795
- sigma: 0.5
- rho: 1.0
- std: None
- 1/tau - sigma||L||^2 >= beta/2: True
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x795b2cd0d690> - 4
- max iterations: 100
- number of reweights: 0
- primal variable shape: (32, 512, 512)
- dual variable shape: (32, 291721)
----------------------------------------
Starting optimization...
WARNING: <class 'mri.operators.linear.wavelet.WaveletN'> does not inherit an operator parent.
- final iteration number: 100
- final cost value: 1000000.0
- converged: False
Done.
Execution time: 208.23918892000074 seconds
----------------------------------------