In this tutorial we will reconstruct an MRI image from Cartesian undersampled kspace data.
Let us denote the undersampling mask, the under-sampled Fourier transform now reads . We use the toy datasets available in pysap, more specifically a 2D brain slice and under-sampled Cartesian acquisition over 32 channels. We compare zero-order image reconstruction with self-calibrated multi-coil Compressed sensing reconstructions (analysis vs synthesis formulation) using the FISTA algorithm for the synthesis formulation and the Condat-Vu algorithm for the analysis formulation. The multicoil data is collected across multiple, say , channels. The sensitivity maps are automically calibrated from the central portion of k-space (e.g. 5%) for all channels . We remind that the synthesis formulation of the non-Cartesian CS-PMRI problem reads (minimization in the sparsifying domain):
and the image solution is given by . For an orthonormal wavelet transform, we have while for a frame we may have . while the analysis formulation consists in minimizing the following cost function (min. in the image domain):
Author: Chaithya G R & Philippe Ciuciu
Date: 01/06/2021, update: 02/13/2024
Target: ATSI MSc students, Paris-Saclay University
#DISPLAY BRAIN PHANTOM
%matplotlib inline
# Package import
from mri.operators import FFT, WaveletN
from mri.operators.utils import convert_mask_to_locations
from mri.reconstructors.utils.extract_sensitivity_maps \
import get_Smaps, extract_k_space_center_and_locations
from mri.reconstructors import SelfCalibrationReconstructor
from pysap.data import get_sample_data
# Third party import
from modopt.math.metrics import ssim
from modopt.opt.linear import Identity
from modopt.opt.proximity import SparseThreshold
import numpy as np
import matplotlib.pyplot as pltLoading input data¶
# Loading input data
cartesian_ref_image = get_sample_data('2d-pmri')
image = np.linalg.norm(cartesian_ref_image, axis=0)
# Obtain MRI cartesian mask
mask = get_sample_data("cartesian-mri-mask").data
# View Input
plt.subplot(1, 2, 1)
plt.imshow(np.abs(image), cmap='gray')
plt.title("MRI Data")
plt.subplot(1, 2, 2)
plt.imshow(mask, cmap='gray')
plt.title("K-space Sampling Mask")
plt.show()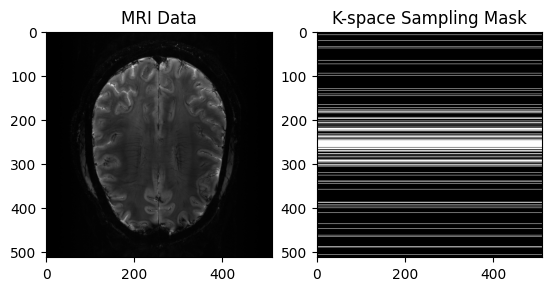
mask_sampled = np.where(mask==1)
print(512**2/np.size(mask_sampled))2.6122448979591835
# Alternative design of 1D VDS
from mrinufft.trajectories.tools import get_random_loc_1d
phase_encoding_locs = get_random_loc_1d(image.shape[0], accel=8, center_prop=0.1, pdf='gaussian')
print(phase_encoding_locs, min(phase_encoding_locs), max(phase_encoding_locs))
phase_encoding_locs = ((phase_encoding_locs +0.5) * image.shape[0]).astype(int)
mask = np.zeros(image.shape, dtype=bool)
mask[phase_encoding_locs] = 1
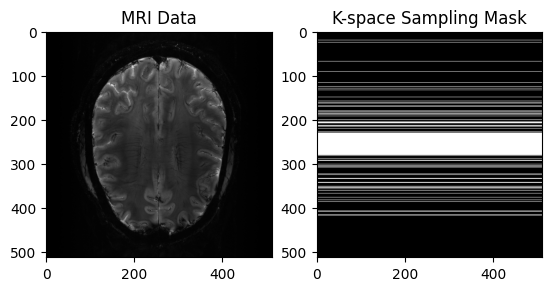
# View Input
plt.subplot(1, 2, 1)
plt.imshow(np.abs(image), cmap='gray')
plt.title("MRI Data")
plt.subplot(1, 2, 2)
plt.imshow(mask, cmap='gray')
plt.title("K-space Sampling Mask")
plt.show()[ 0. 0.00195312 -0.00195312 0.00390625 -0.00390625 0.00585938
-0.00585938 0.0078125 -0.0078125 0.00976562 -0.00976562 0.01171875
-0.01171875 0.01367188 -0.01367188 0.015625 -0.015625 0.01757812
-0.01757812 0.01953125 -0.01953125 0.02148438 -0.02148438 0.0234375
-0.0234375 0.02539062 -0.02539062 0.02734375 -0.02734375 0.02929688
-0.02929688 0.03125 -0.03125 0.03320312 -0.03320312 0.03515625
-0.03515625 0.03710938 -0.03710938 0.0390625 -0.0390625 0.04101562
-0.04101562 0.04296875 -0.04296875 0.04492188 -0.04492188 0.046875
-0.046875 0.05078125 -0.04882812 0.0625 -0.05078125 0.0703125
-0.05664062 0.07226562 -0.0703125 0.08789062 -0.078125 0.09375
-0.08203125 0.09765625 -0.08398438 0.1015625 -0.09375 0.13085938
-0.09570312 0.13476562 -0.09765625 0.15039062 -0.1015625 0.15234375
-0.109375 0.16796875 -0.125 0.16992188 -0.1328125 0.18554688
-0.13476562 0.18945312 -0.13867188 0.19140625 -0.14453125 0.19726562
-0.1484375 0.20507812 -0.16796875 0.21679688 -0.16992188 0.234375
-0.18164062 0.24609375 -0.18554688 0.25390625 -0.19140625 0.29492188
-0.20898438 0.296875 -0.24023438 0.31054688 -0.24804688 0.31445312
-0.25585938 -0.2734375 -0.32226562 -0.3671875 -0.45117188 -0.4609375 ] -0.4609375 0.314453125
Generate the kspace¶
From the 2D brain slice and the acquisition mask, we retrospectively undersample the k-space using a cartesian acquisition mask. We then reconstruct the zero order solution as a baseline
# Get the locations of the kspace samples and the associated observations
fourier_op = FFT(mask=mask, shape=image.shape,
n_coils=cartesian_ref_image.shape[0])
kspace_obs = fourier_op.op(cartesian_ref_image)# Zero order solution
zero_soln = np.linalg.norm(fourier_op.adj_op(kspace_obs), axis=0)
base_ssim = ssim(zero_soln, image)
plt.imshow(np.abs(zero_soln), cmap='gray')
plt.title('Zero Order Solution : SSIM = ' + str(np.around(base_ssim, 3)))
plt.show()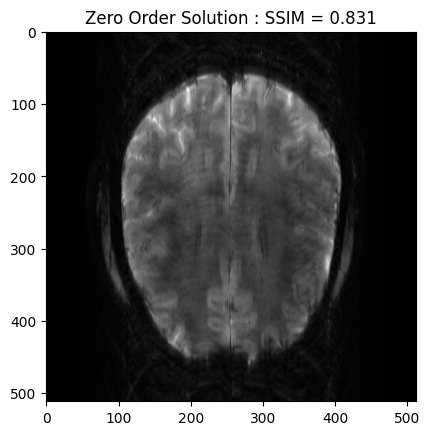
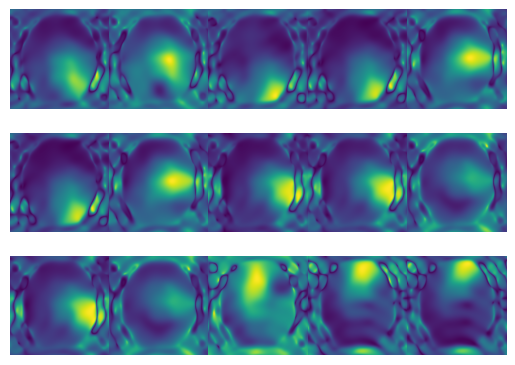
# Obtain SMaps
kspace_loc = convert_mask_to_locations(mask)
Smaps, SOS = get_Smaps(
k_space=kspace_obs,
img_shape=fourier_op.shape,
samples=kspace_loc,
thresh=(0.01, 0.01), # The cutoff threshold in each kspace direction
# between 0 and kspace_max (0.5)
min_samples=kspace_loc.min(axis=0),
max_samples=kspace_loc.max(axis=0),
mode='gridding',
method='linear',
n_cpu=-1,
)
h=3;w=5;
f, axs = plt.subplots(h,w)
for i in range(h):
for j in range(w):
axs[i, j].imshow(np.abs(Smaps[3 * i + j]))
axs[i, j].axis('off')
plt.subplots_adjust(wspace=0,hspace=0)
plt.show()[Parallel(n_jobs=-1)]: Using backend LokyBackend with 20 concurrent workers.
[Parallel(n_jobs=-1)]: Done 26 out of 32 | elapsed: 0.2s remaining: 0.1s
[Parallel(n_jobs=-1)]: Done 32 out of 32 | elapsed: 0.3s finished
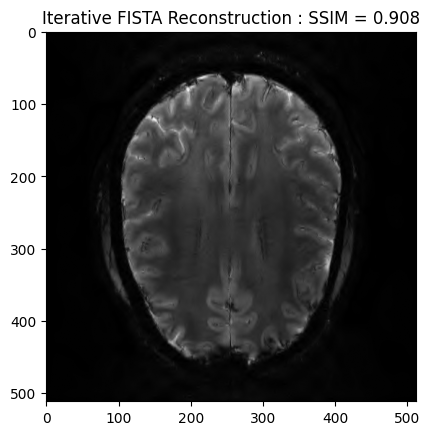
FISTA optimization¶
We now want to refine the zero order solution using a FISTA optimization. The cost function is set to Proximity Cost + Gradient Cost
# Setup the operators
linear_op = WaveletN(
wavelet_name='sym8',
nb_scale=4,
)
regularizer_op = SparseThreshold(Identity(), 1.5e-8, thresh_type="soft")# Setup Reconstructor
reconstructor = SelfCalibrationReconstructor(
fourier_op=fourier_op,
linear_op=linear_op,
regularizer_op=regularizer_op,
gradient_formulation='synthesis',
kspace_portion=0.01,
verbose=1,
)image_rec, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='fista',
num_iterations=200,
)
recon_ssim = ssim(image_rec, image)
plt.imshow(np.abs(image_rec), cmap='gray')
plt.title('Iterative FISTA Reconstruction : SSIM = ' + str(np.around(recon_ssim, 3)))
plt.show()WARNING: Making input data immutable.
Lipschitz constant is 1.0983724828872625
The lipschitz constraint is satisfied
- mu: 1.5e-08
- lipschitz constant: 1.0983724828872625
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x776fb38239d0> - 4
- max iterations: 200
- image variable shape: (512, 512)
- alpha variable shape: (291721,)
----------------------------------------
Starting optimization...
WARNING: Making input data immutable.
- final iteration number: 200
- final log10 cost value: 6.0
- converged: False
Done.
Execution time: 98.67630440200446 seconds
----------------------------------------
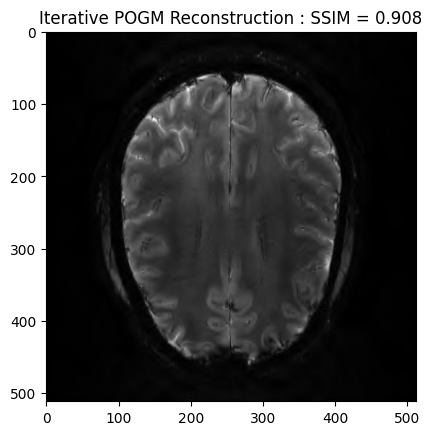
POGM reconstruction¶
image_rec2, costs, metrics = reconstructor.reconstruct(
kspace_data=kspace_obs,
optimization_alg='fista',
num_iterations=200,
)
recon2_ssim = ssim(image_rec2, image)
plt.imshow(np.abs(image_rec2), cmap='gray')
plt.title('Iterative POGM Reconstruction : SSIM = ' + str(np.around(recon2_ssim, 3)))
plt.show()WARNING: Making input data immutable.
Lipschitz constant is 1.09836964369987
The lipschitz constraint is satisfied
- mu: 1.5e-08
- lipschitz constant: 1.09836964369987
- data: (512, 512)
- wavelet: <mri.operators.linear.wavelet.WaveletN object at 0x776fb38239d0> - 4
- max iterations: 200
- image variable shape: (512, 512)
- alpha variable shape: (291721,)
----------------------------------------
Starting optimization...
- final iteration number: 200
- final log10 cost value: 6.0
- converged: False
Done.
Execution time: 101.1655332060036 seconds
----------------------------------------